Sala de prensa
Actualidad
Todas las noticias
-

-
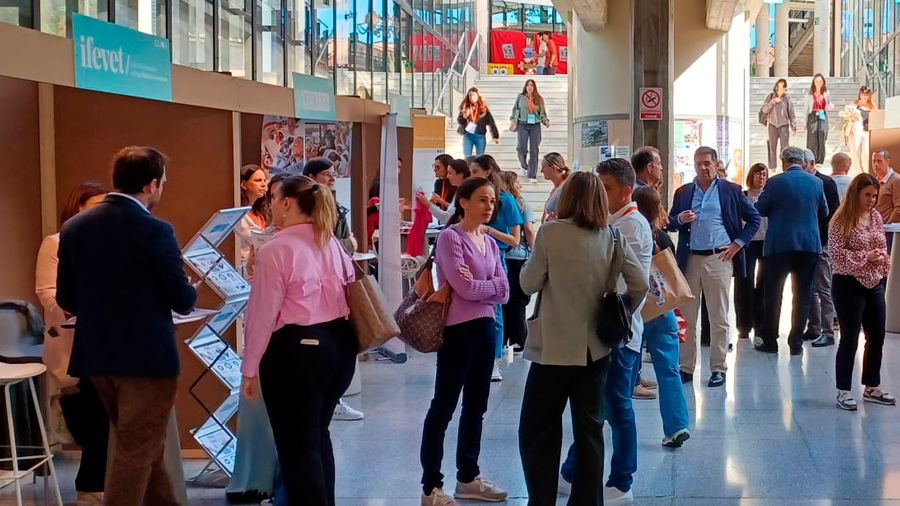

Éxito de la primera Jornada Ocupacional en Veterinaria
El jueves 18 de abril ha tenido lugar, con gran éxito de participación, la primera Jornada Ocupacional en la Facultad de Veterinaria, organizada para dar...
-

Abierta una nueva convocatoria de profesorado sustituto para el curso 2024-2025
La UAB abre la tercera convocatoria para confeccionar las bolsas de profesorado sustituto por cada una de las áreas de conocimiento. El plazo de presentación...
-
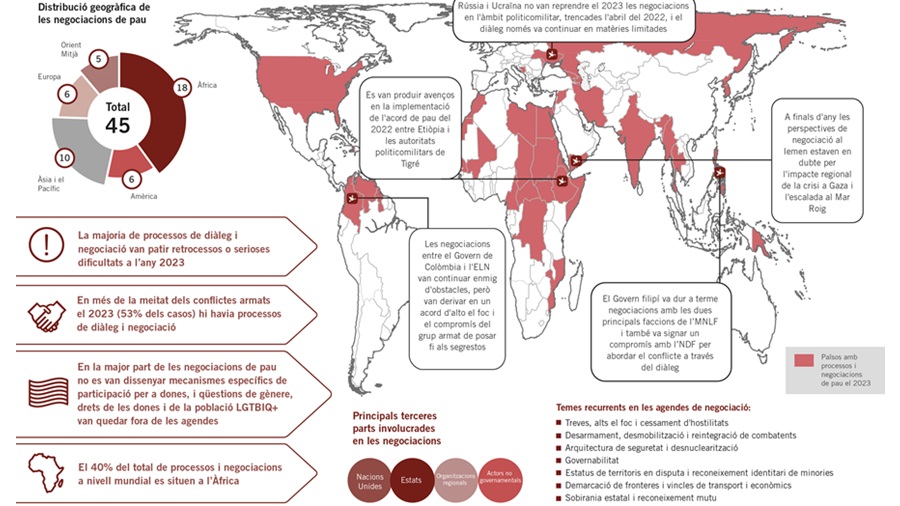
Durante 2023 se identificaron 45 procesos y negociaciones de paz en el mundo
La Escuela de Cultura de Paz de la UAB ha publicado un año más su anuario Negociaciones de paz 2023. Análisis de tendencias y escenarios, en el que recoge...
-
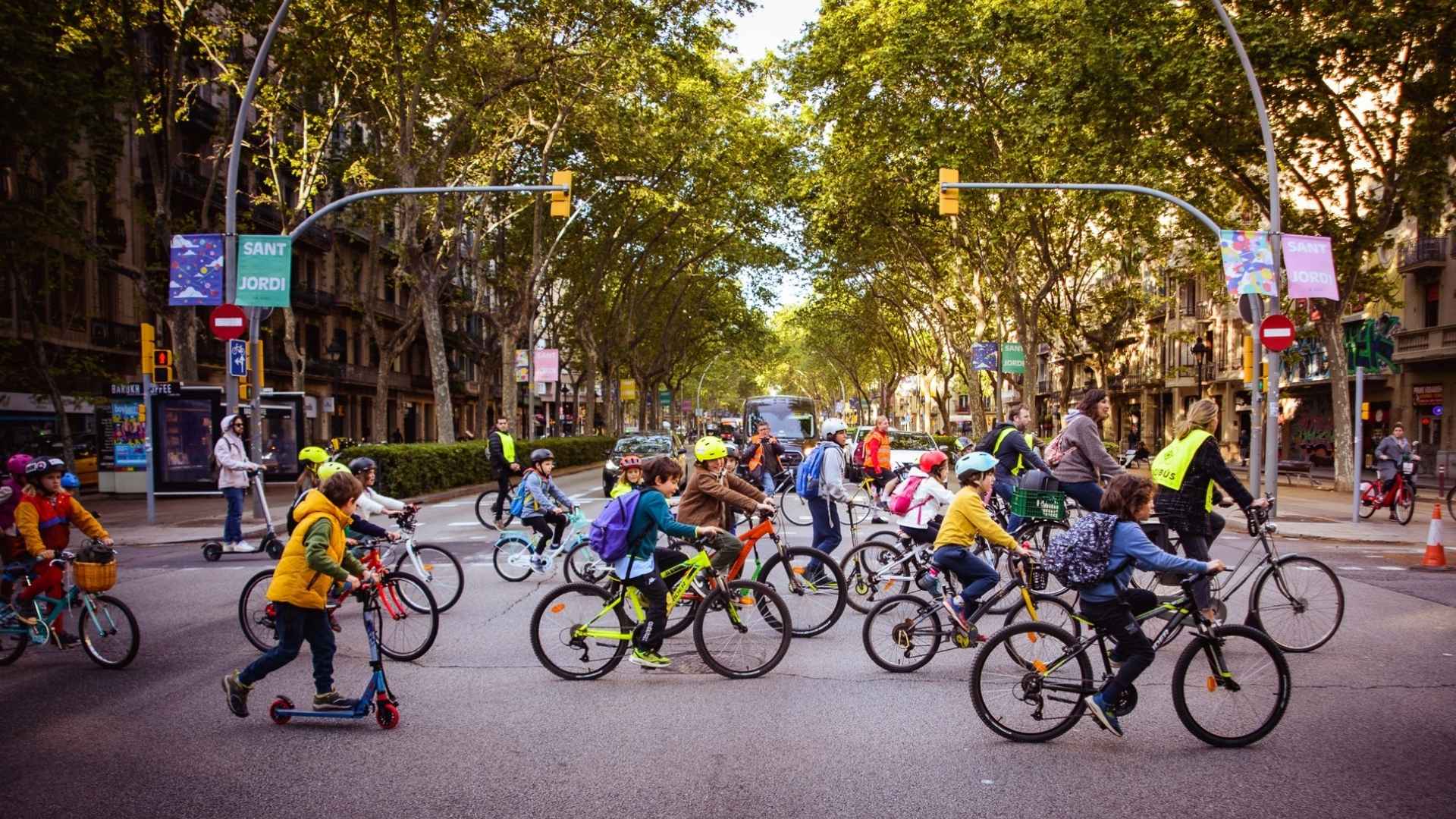

El bicibús gana adeptos como una forma de reivindicación de la movilidad sostenible y segura
En un mundo cada vez más consciente del cambio climático y la importancia del transporte activo, el movimiento Bicibús ha surgido como una poderosa...
-

Entregados los premios a los mejores TFG de transformación para la justicia global
La Fundació Autònoma Solidària (FAS) ha entregado los premios a los mejores trabajos de fin de grado (TFG) de transformación para la justicia global —que...
-
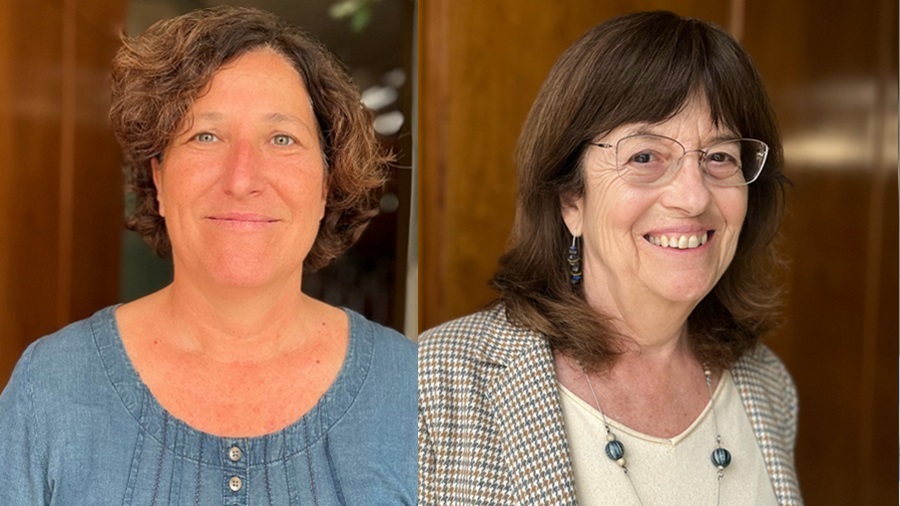

Reestructuración del Equipo de Gobierno de la UAB
Tatiana Rovira es la nueva vicerrectora de Estudios y de Calidad y Laura Santamaria es la nueva vicerrectora de Relaciones Internacionales y de Política...
-

Jornada La educación democrática y en derechos humanos - 18 de abril
El próximo 18 de abril DIPLOCAT y CATESCO organizan una jornada sobre La educación democrática y en derechos humanos para contrarrestar el discurso de odio.
-

Nueva edición del concurso fotográfico EXPONATURA UAB
El Departamento de Biología Animal, de Biología Vegetal y de Ecología organiza una nueva edición del concurso EXPONATURA UAB, el concurso fotográfico en...
-
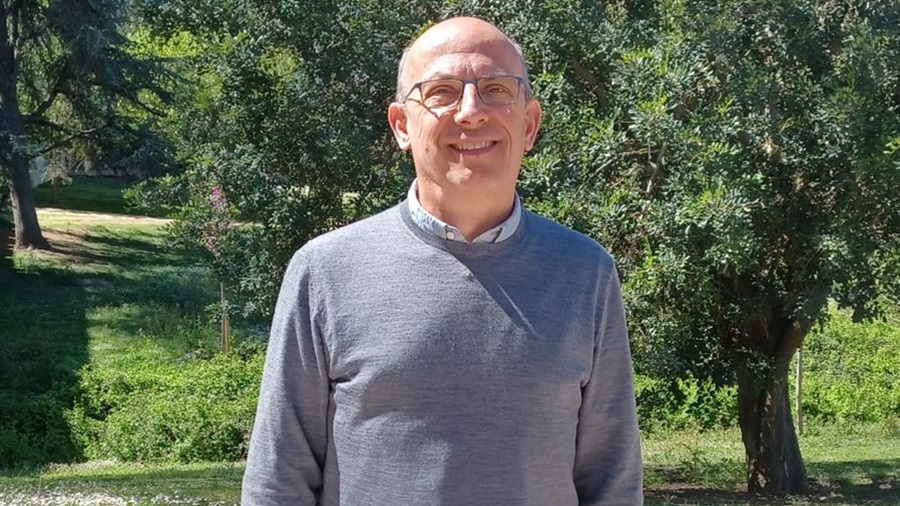

Xavier Ballart, nuevo decano de la Facultad de Ciencias Políticas y de Sociología
Xavier Ballart, catedrático del Departamento de Ciencia Política y Derecho Público de la UAB, ha sido elegido nuevo decano de la Facultad de Ciencias...
Ver noticias de los últimos 30 días
Para buscar otra franja de fechas, categorías o palabras clave se puede utilizar el filtro situado en la parte superior de la página.